 Image via Wikipedia
Image via WikipediaPerkins Elmer has an extensive history in the genomic sequencing field and Dr. Craig Venter. Dr. Venter is a former researcher for The Institute for Genomic Research and founded the J. Craig Venter Institute. Find out more here: http://www.jcvi.org/
Google finance: NYSE:PKI
PerkinElmer Website
PerkinElmer Enters DNA Sequencing Market
By MATTHEW HERPER
PerkinElmer headquarters
In a move that heralds the growing importance of DNA sequencing to medical research and, eventually, diagnostics, PerkinElmer, the $1.9 billion (sales) maker of diagnostic tests, is entering the gene scanning market.
Rather than compete with makers of big DNA sequencing machines, PerkinElmer is creating a service business that will allow researchers to get human genetic data without owning their own DNA sequencers or high-powered supercomputers. Customers will be able to access and analyze DNA data on a password-protected computer cloud. The DNA sequencing and the data analysis will be compliant with regulations for lab diagnostics. Eventually, the will also comply with the Health Insurance Portability and Accountability act (HIPAA), which protects medical privacy.
Continued
The decision to enter the DNA-decoding market came about because although PerkinElmer is big player in diagnostics and in the analysis of proteins, it had no real presence in DNA sequencing.”You can’t really develop drugs without spending much more energy on the DNA side,” says Richard Begley, president of emerging technologies at Perkin, in an exclusive interview. “The absence of DNA was clearly a problem.”
 Image via Wikipedia
Image via WikipediaThere’s a historic resonance to the move, too. The early DNA sequencers that were used in the Human Genome Project started out at Perkin-Elmer before a restructuring made the technology part of a separate company. Begley, the executive heading the DNA sequencing effort, was previously chief executive of 454 Life Sciences, the first of the current generation of faster, more powerful DNA decoders.
DNA sequencing has become a booming growth market. The machines and the chemicals that run them are a $1.5 billion market dominated by Illumina, Life Technologies, and Roche, which bought 454 in 2007. Their ascent has made Illumina one of the hottest stocks of the past five years as its sales have grown 12-fold to a projected $880 million in 2010.
PerkinElmer will focus its product on what is known as exome sequencing, a technique for looking at only the parts of the human genome that contain genes that code for proteins. (This is only 1% of the 6 billion DNA bases in a human genome; the rest either regulates the genes or is junk.) This year, researchers are expected to sequence thousands of exomes in order to understand rare diseases and cancers.
Begley says that PerkinElmer already has 20 customers for its sequencing service and that it has already delivered hundreds of exomes. The company was not able to provide any customers to be interviewed for this story.
This is hardly the first sequencing-as-service business. Complete Genomics has been in the service business for years, with clients that have included Pfizer, Eli Lilly, and the Institute for Systems Biology, and it has completed hundreds of full human genomes. Illumina has its own sequencing business, as does Knome of Cambridge, Mass. The Beijing Genomics Institute does some sequencing for other researchers. Complete has referenced per-genome prices of as low as $5,000.
 Image by Getty Images via @daylife
Image by Getty Images via @daylifeThe PerkinElmer solution will allow customers to choose between technologies made by Roche, Illumina, and Agilent to cut genomes down to exome size, but all the sequencing will be done on machines made by Illumina. In the future, Begley says, PerkinElmer will use whichever sequencing platform meets its needs or is wanted by enough customers.
The systems are automated using PerkinElmer technology and software is provided by Geospiza, a computing firm. Begley declined to provide a per exome price estimate for the new service, but said that it was competitive with other players. For more information, go here.
Jan. 24 2011 — 8:09 am
Perkin-Elmer, Dr. Craig Venter, and TIGR Announce Formation of New Genomics Company
-- Plan to Sequence Human Genome Within Three Years --
May 9, 1998
09-May-1998
PRESS RELEASE
 Image via Wikipedia
Image via WikipediaNORWALK, CT and ROCKVILLE, MD, May 9, 1998 -- The Perkin-Elmer Corporation (NYSE:PKN), Dr. J. Craig Venter, and The Institute for Genomic Research (TIGR) announced today that they have signed letters of intent relating to the formation by Perkin-Elmer and Dr. Venter of a new genomics company. Its strategy will be centered on a plan to substantially complete the sequencing of the human genome in three years.
The new company's goal is to become the definitive source of genomic and associated medical information that will be used by scientists to develop a better understanding of the biological processes in humans and deliver improved healthcare in the future. Using breakthrough DNA analysis technology being developed by Perkin-Elmer's Applied Biosystems Division, applied to sequencing strategies pioneered by Dr. Venter and others at TIGR, the company will operate a genomics sequencing facility with an expected capacity greater than that of the current combined world output.
Concurrently, the new company also intends to build the scientific expertise and informatics tools necessary to extract valuable biological knowledge from genomic data, including the discovery of new genes, development of polymorphism assay systems, and databases for the scientific community. Perkin-Elmer and Dr. Venter believe that this information has significant commercial value and that the new company can provide this information more rapidly and more accurately than is currently possible.
 Image via Wikipedia
Image via Wikipedia"The fundamentals of healthcare delivery and medical practice will be transformed by molecular medicine. We believe that the information developed by this new company will become the cornerstone of this new paradigm and accelerate development of new therapies, targeted diagnostics, and individualized medicine - providing therapies tailored specifically to a disease in an individual," noted Tony L. White, Perkin-Elmer's chairman, president and chief executive officer.
Mr. White continued, "We see our exciting new sequencing technology as the catalyst for this new genomics initiative - allowing us to take another step in the redefinition of Perkin-Elmer. When combined with the talents and resources of Dr. Venter and other TIGR scientists who would be employed by the new company, we believe we have the ability to quickly and cost-effectively sequence whole genomes. Together, we intend to build a new business that satisfies the rapidly growing need for genomic information and gene discovery."
Since the inception of the Human Genome Project (HGP) in 1990, a major shift in technology has been anticipated that would allow the entire sequence to be completed. To date, scientists have constructed detailed genetic and physical maps and sequenced approximately three percent of the three billion base pairs of DNA that comprise the human genome and contain the inherited instructions for human development and function.
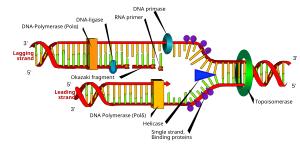 Image via Wikipedia
Image via Wikipedia"The Human Genome Project has been a technology-driven quest," said Dr. Michael W. Hunkapiller, senior vice president of Perkin-Elmer and president of its Applied Biosystems Division. "We're near the end of our technology development phase and about to implement a new sequencing strategy. We believe we can provide the anticipated advance that greatly expedites the sequencing phase of the entire human genome."
The new company plans to make sequencing data publicly available to ensure that as many researchers as possible are examining it and that applications, such as the development of diagnostic tests and new drug discovery, are as broad as possible.
Commenting on the proposed new venture, J. Craig Venter, Ph.D., TIGR's president and director, said, "By linking techniques that have been used by TIGR scientists with Perkin-Elmer's genetic analysis technologies, we pave the way for a new era of post-genomic discovery. The sooner researchers can access the information contained in the complete human genome, the sooner new therapies can be developed for the thousands of disorders in which genes play a role."
Dr. Venter will serve as the president of the proposed new company. Mr. White will serve as the new firm's chairman, and Perkin-Elmer's vice president of corporate planning and business development, Peter Barrett, Ph.D., will be appointed an executive vice president. Perkin-Elmer will retain ownership of approximately 80 percent of the new company, with the balance being held by TIGR, Dr. Venter, other members of management, and employees of the new company in the form of common shares and options. The company will be located in Rockville, MD.
Additional terms of the letters of intent were not disclosed, and the agreement is subject to final documentation and the approval of the boards of Perkin-Elmer and TIGR.
The Perkin-Elmer Corporation is a leading supplier of systems for life science research and related applications. It develops, manufactures, and markets life science systems and analytical instruments used in markets such as pharmaceutical, biotechnology, environmental testing, foods, agriculture, and chemical manufacturing. Headquartered in Connecticut, Perkin-Elmer had revenues of nearly $1.4 billion in fiscal 1997 and employs more than 6,000 people worldwide.
TIGR, headquartered in Rockville, MD is an independent, not-for-profit research institute founded in 1992 by Dr. Venter that employs 170 faculty and staff. TIGR has interest in structural, functional and comparative analysis of genome and gene products in viruses, eubacteria, pathogenic bacteria, archaea, and eukaryotes, both plant and animal, including humans. In its brief history, TIGR has fully sequenced seven organisms. Most recently, TIGR released the DNA sequence for H. pylori, the bacteria that causes stomach ulcers and B. burgdorferi, the pathogen that causes Lyme disease.
Perkin-Elmer will hold a conference call, at 10:00 a.m. ET on Monday, May 11, to discuss this press release. To participate in this call, those interested should phone (203) 761-2617 to receive the conference number. This and other information about the Company is also available on the World Wide Web at www.perkin-elmer.com or by phoning (800) 762-6923.
 Image via Wikipedia
Image via WikipediaSequencing Technologies
The Scientist 1995, 9(20):18
Published 16 October 1995
--------------------------------------------------------------------------------
Genome
Author: Holly Ahern
SIDEBAR: Selected Suppliers of DNA Sequencing Equipment and Supplies
Since the Human Genome Project (HGP) was launched five years ago, human geneticists working to decipher the code of nucleotides in the DNA of human cells have enthralled the public with discoveries of numerous genes that are responsible for human diseases, such as cancer-related genes. Different groups of scientists in laboratories all over the world are participating in this project, taking apart the human genome right down to the constituent DNA nucleotides. The ultimate goal of the HGP is to determine the exact order, or DNA sequence, of the nucleotide bases that make up all 23 pairs of human chromosomes, some 3 billion bases in all.
As high-profile as the project may be, HGP researchers aren't the only molecular geneticists who are attracting fame and glory. In July, a research team headed by Nobel laureate Hamilton Smith of Johns Hopkins University and Craig Venter of The Institute for Genomic Research in Gaithersburg, Md., published the first complete genomic sequence of the bacterium Haemophilus influenzae, a laboratory organism and potential pathogen. H. influenzae gained the "honor" of becoming the first free-living organism whose genetic composition is completely known. (R.D. Fleischmann et al., Science, 269:496-512, 1995).
"This is a notable effort," comments Barton Slatko, director of the DNA sequencing core facility at New England BioLabs Inc. in Beverly, Mass. "The Human Genome Project gains a lot of attention, but this shows that there are many other applications of DNA sequencing technology that are just as noteworthy."
SEQUENCING BY CAPILLARY ELECTROPHORESIS: the ABI Prism 310 Genetic Analyzer from Perkin-Elmer Biosystems is a sequencing and DNA-fragment znalyzing system using capillary electrophoresis.
 Image via Wikipedia
Image via WikipediaThe implications of Venter and Smith's research are far- reaching. With the sequence data generated from this effort and from sequencing the genomes of other microbes and eukaryotes, researchers might find answers to the question of how microorganisms and higher creatures evolved. Additionally, investigators sorting through the 1,743 genes in H. influenzae's genome have found genes of unknown function as well as counterparts to known genes in other organisms. Knowing the identity of all of H. influenzae's genes will also aid medical microbiologists studying the organism's natural pathogenicity.
New sequencing methods, along with automated instruments and computerized analysis of the sequencing data, are the tools that researchers such as Venter and Smith are using to generate genomic sequences in a fraction of the time it takes to achieve the same results manually. The techniques used by Venter's group to sequence the bacterial genome may have direct bearing on the HGP if their methods are applicable to pieces of human DNA. Because Venter and Smith's method involves less initial cloning, sequencing time -- and, thus, the cost -- are greatly reduced.
With conventional methods, in order to determine the nucleotide sequence of something as big as an organism's full contingent of DNA, researchers first must painstakingly clone the genomic DNA piece by piece in large (approximately 40 kilobases), overlapping segments. Each segment is "shotgunned," or broken into smaller fragments, which serve as templates for sequencing. The order of bases in each fragment is determined with a series of chemical reactions followed by electrophoresis to separate the resulting DNA pieces, which differ in length by only one base. Once the order is determined, the shotgunned fragments are ordered according to overlaps in their sequence. Then the larger segments are put back into place in the genome using the same type of analysis -- not a trivial accomplishment, and one that requires powerful computing capabilities.
This approach was used successfully by Venter's team. They shotgunned the entire 1,800-kilobase H. influenzae genome, sequenced each fragment, and then reassembled the genome by lining up overlapping ends. To reassemble the genome from the tens of thousands of fragments generated by shotgunning, the research group relied on their own program, called TIGR Assembler, to put the genome back together again. Although the human genome is roughly 1,500 times larger than that of the bacterium, the researchers are optimistic that this new method can be adapted for sequencing human DNA.
When the HGP began in 1990, sequencing was a manual effort that was both labor-intensive and expensive, which prompted former HGP director James D. Watson and his associates to propose that the genome be divvied up among several laboratories all over the world. Manually sequencing even a small fragment of DNA is a complicated process in itself, requiring special electrophoresis equipment and an extensive array of reagents and chemicals for purifying DNA, pouring gels, and performing the sequencing reactions. Talent and patience are also necessary. Pouring a good sequencing gel, which is a very thin slab of acrylamide poured between two long glass plates, can be more of an art than a science.
 Image via Wikipedia
Image via Wikipedia"In general, pouring gels is something that many people find technically difficult," notes Randall Collura, a molecular biologist at the State University of New York, Albany. Collura runs a DNA sequencing facility for members of the molecular biology core group at the university. "With all of the available kits and enzymes, conventional sequencing chemistry is quite straightforward. but the quality of the data really depends on how well the DNA fragments separate in the gel," he says.
Anomalies commonly encountered with sequencing gels, such as compressed bands (bands that are squeezed together) and "smiling" gels (occurring when genes at the beginning and end of the band travel more slowly through the gel than the ones in the middle, causing the gel to look like a smile), make it difficult to read long stretches of sequence. Investigators may be able to read only 100 bases rather than the 300 or 400 they would normally read from a very good gel. To generate further sequence data, more cloning and sequencing steps must be performed. This translates into lost time and money.
Advances in sequencing technology stemming from HGP research have led to improvements in gel polymers, gel pouring techniques, and the requisite electrophoresis equipment. The gel polymer most often used in sequencing, polyacrylamide, can be purchased from companies such as Bio-Rad Laboratories in Hercules, Calif., which offers several different formulations of acrylamide powders and solutions, and Pharmacia Biotech Inc. of Piscataway, N.J. Pharmacia's PlusOne acrylamide products offer researchers a range of acrylamide mixtures and come as powders or as solutions, depending on the investigator's preference.
Electrophoresis "rigs" for sequencing -- which include gel assembly and support components, glass plates, combs for sample loading, and a power supply -- are improving, as well. Many electrophoresis units come complete with temperature controls that allow investigators to regulate the temperature of the gel, which generally goes way up as the applied voltage is increased. With this feature, gels can be run faster without worries of "cooking" the gel, or ending up with smiles or compressed bands.
Electrophoresis instruments, such as Pharmacia's Macrophor system, feature a thermostatic plate that keeps the entire gel at a uniform temperature. Using the MacroMould Gel Casting Unit (also from Pharmacia), researchers create ultrathin gels (100 mm thick) that can be cast in less than one minute. Bio-Rad's Sequi-Gen II Sequencing Cell incorporates an upper buffer chamber that extends over the entire length of the gel to dissipate heat and maintain the gel at a constant temperature.
 Image via Wikipedia
Image via WikipediaThe Model SA Sequencing Gel Electrophoresis System from Life Technologies Inc. of Gaithersburg, Md., is an adjustable system for gels of different heights. For sequencing reactions, for example, a researcher would choose the longest gel configuration (88 cm). The system also includes an integral aluminum plate design that distributes heat evenly across the gel to minimize electrophoretic anomalies.
"With DNA sequencing, there are essentially five areas of consideration," explains New england BioLab's Slatko. The first of these considerations are the DNA-based steps of cloning to mass-produce the area of DNA to be sequenced and purifying the target to be sequenced. For example, the targets may be templates such as single-stranded or double- stranded DNA, messenger RNA, or polymerase chain reaction products. Researchers must also choose both the type of sequencing chemistry (Maxam-Gilbert, Sanger, or thermal cycling) and an isotopic or nonradioactive labeling and detection system. A fourth consideration, for separating the fragments generated by the sequencing reactions, is the electrophoresis rig and the method of gel process, usually autoradiography or gel-scanning devices that read bands directly from an electrophoresis gel. The final consideration is the interpretation of gel patterns to obtain the DNA-sequence information, which is usually performed with the aid of a computer and dedicated software.
Each of these areas has seen advancements, many of which are related to the HGP and other genome-sequencing efforts. "The Human Genome Project has dedicated large amounts of money and resources for developing new and advanced sequencing technologies," says Collura. Research aimed at developing techniques that will decrease the length of the project as well as project costs is generally well funded. "Any improvement that makes sequencing faster, cheaper and easier to perform will pay off in the end," Collura notes.
Cycle-sequencing chemistry, sequencing based on DNA amplification, is an example. Most conventional sequencing strategies are based on the dideoxy chain-termination method developed by Fred Sanger (F. Sanger et al., Proceedings of the National Academy of Sciences, 74:5463, 1977). It uses a radioactive isotope as the label to create visible bands on an X-ray film after electrophoresis of the chain-terminated DNA fragments. Cycle sequencing combines conventional dideoxy-sequencing reactions with thermal-cycling conditions, and involves a thermostable DNA polymerase, such as Taq, and a thermal cycler for repeated rounds of DNA denaturation, annealing, and primer extension. This offers many advantages over a single-cycle method, because the reactions can be performed at higher temperatures (preventing false-priming reactions) and less template is required, since the product is linearly amplified over several cycles. Bacterial colonies or phage plaques, as opposed to highly purified DNA, can be used as template in this type of reaction.
Several cycle-sequencing kits that contain all of the necessary reactants (a thermal-cycling instrument is also required) are commercially available. The Cycle Sequencing Kit from Pharmacia generates approximately 500 bases of readable sequence data from as little as 50 femtomoles (10- 15 moles) of DNA. The Cyclist Taq DNA Sequencing kit from Stratagene Cloning Systems in La Jolla, Calif., is similarly based on thermal cycle-sequencing chemistry.
New England BioLabs uses a highly thermostable polymerase (Vent exo- DNA polymerase) in its CircumVent Thermal Cycle DNA Sequencing kit. Combining this product with the CircumVent Phototope Detection kit for chemiluminescent detection of DNA bands, researchers can sequence DNA without radioactivity.
For scientists who prefer the nonradioactive sequencing strategies, Tropix Inc. of Bedford, Mass., markets a chemiluminescence-based DNA sequencing product, the SEQ- Light DNA-sequencing system, that combines standard dideoxy- sequencing chemistry with chemiluminescent detection. Boehringer Mannheim Corp. of Indianapolis also carries a sequencing kit for those who prefer the nonradioactive approach. The Genius Nonradioactive DNA Sequencing kit can be used for both standard or cycle-sequencing with chemiluminescent detection.
Most of the recent sequencing developments are aimed at speeding up the sequencing process and increasing the length of "read," or how many bases in a row a researcher can accurately read after a single electrophoretic run. The most direct way to achieve these goals is to automate some or all of the sequencing steps.
Preparing templates for sequencing, for example, can be automated with the Vistra DNA Labstation 625, manufactured by Molecular Dynamics Inc. of Sunnyvale, Calif. The unit reportedly performs all of the steps required to prepare samples for sequencing-gel loading. The starting material can be purified DNA, or simply the cells or virus particles containing the target template. For loading gels, Bio-Rad carries an automatic gel loader (GS Gene Loader II) that adds radiolabeled or fluorescently labeled samples onto sequencing gels in semiautomated sequencing systems, such as those offered by Perkin-Elmer Corp. Applied Biosystems Division in Foster City, Calif., and Pharmacia.
The GS Gene Reader from Bio-Rad is an automated film scanner that simplifies one of the most tedious sequencing steps -- reading the order of bases. When connected to a computer workstation, the scanning system delivers more bases in less time and allows the researcher to process images, assign bases, and edit sequence data. To read sequence data directly from gels, Molecular Dynamics markets two systems: the FluorImager SI, for fluorescent labels; and the PhosphorImager 445 SI, which captures images produced by radioactive emissions from isotopes. Molecular Dynamics also carries software (DNAscan) for computer-assisted DNA sequencing of images from films and gels.
Companies such as Molecular Dynamics; Genomyx Corp. of Foster City, Calif.; Perkin-Elmer Applied Biosystems; and Pharmacia market semiautomated sequencing instruments that reportedly sequence more bases in less time and at a fraction of the cost of manual sequencing (according to the company, as low as 5 to 10 cents a base). Additionally, Hyseq Inc. of Sunnyvale, Calif., is currently developing -- but has not yet made available -- its sequencing-by-hybridization (SBH) technology, whereby sequencing is performed directly on a microchip. The chip serves as a physical platform where target sequences hybridize with immobilized oligonucleotides (eight bases long). Computerized analysis of the hybrids then provides the researcher with the unknown sequence, at a rate of up to 64,000 bases of DNA in less than two hours, according to the company.
Speedy sequences don't come cheap, however. You have to buy the instrument first, a one-time investment of $10,000 to $100,000 or more, depending on the instrument.
ALL IN ONE: The Genomyx genomyxLR DNA Sequencer replaces the conventional electrophoresis "rig", including the power supply and gel drier.
 Image via Wikipedia
Image via Wikipedia--------------------------------------------------------------------------------
The genomyxLR DNA Sequencer from Genomyx Corp., priced slightly less than $10,000, is a completely integrated system that replaces the conventional electrophoresis "rig," including the power supply and gel drier (along with the associated cold trap and vacuum pump). This system, which is semiautomated (investigators pour and load gels, as with manual methods), features air-impingement technology that uniformly distributes air over the glass plates, eliminating temperature gradients or differences in the gel. Sequencing gels can be run at higher temperatures or higher voltage with good resolution of the DNA fragments.
The genomyxLR also has a unique incorporated radionucleotide capture cartridge for processing radioactive buffer left over at the end of an electrophoresis run. This feature decreases the amount of liquid radioactive waste to nearly nonexistent, because the radioactivity is collected in the cartridge. Next year Genomyx plans to introduce a fluorescent sequencer as a nonradioactive alternative to conventional sequencing protocols.
Molecular Dynamics' Vistra DNA Sequencer 725 is a mid-range semiautomatic sequencing instrument, priced at $49,500 without a computer workstation and at $54,900 with a workstation.
Pharmacia and Applied Biosystems both carry semiautomated sequencing systems for their higher-end users. Priced at $75,000, Pharmacia's ALF DNA Analysis System (ALF DNA Sequencer) and Pharmacia's newer, less costly system, the ALFexpress, utilize a single fluorescent dye with LIF (laser-induced fluorescence) detection. The ALF system incorporates an electrophoresis module that includes the argon laser for detection, a power module with both electrophoretic and laser power supplies, and a PC-based computer system with system-management software. Three dedicated reagent kits -- the AutoRead Sequencing kit, AutoRead 1000 Sequencing kit, and the AutoCycle Sequencing Kit for thermal cycle sequencing-support the system.
 Image via Wikipedia
Image via WikipediaThe ABI Prism 377 DNA sequencer from Perkin-Elmer Applied Biosystems, which succeeds ABI's model 373 instrument, is a high-throughput DNA sequencer that runs in a slab-gel format (referring to the type of gel, in this case a thin polyacrylamide slab). Like the 373 model, the 377 is based on the company's four-color fluorescent labeling technology. With four fluorescent dyes (for the four bases in DNA), all of the sequencing reactions can be run in one lane. This increases the number of bases that can be read from a single gel and generates sequencing data faster, at a rate of up to 7,200 bases per hour. However, with a price tag of more than $100,000, this instrument is largely relegated to core facilities that support a large number of researchers. Users of the core facility generally pay to have their DNA sequenced, usually at a per-base rate.
Newer to the market is the ABI Prism 310 Genetic Analyzer from Perkin-Elmer Applied Biosystems, a near fully automated sequencing and DNA-fragment analyzing system priced at less than half the cost of the model 377. With the ABI Prism 310, sequences are determined by capillary electrophoresis, whereby the DNA pieces generated in the sequencing reactions are separated in a polymer gel contained within a glass capillary tube. With the exception of purifying the DNA and performing the chemical reactions, the model 310 is fully automated. Prepared samples are injected into the capillary from a sample tray, and the fragments are separated and analyzed by four-color LIF detection.
"Four-color LIF DNA sequencing utilizes four different [fluorescent] color labels, one for each DNA nucleotide," explains J. William Efcavitch, genetic-analysis research and development section leader at Perkin-Elmer. "All four reactions are loaded into the systems' single capillary, where the differently sized fragments are separated" and then analyzed, at a rate of 400 to 450 bases every two hours. "The second generation of this instrument [due out in January] is expected to sequence 600 to 650 nucleotides in the same amount of time," he adds.
 Image via Wikipedia
Image via Wikipedia"When you are sequencing DNA, it's a simple choice," according to Efcavitch. "You can read a fewer number of bases at ultra-high speed, or you can read more bases but at a slower rate." To sequence a genome, investigators are interested in as much throughput as possible. For smaller projects, such as in the applied sciences of forensics and diagnostics, where only bits of genome are examined, Efcavitch concludes, "the greater need is for total automation and longer read, as opposed to faster throughput."
Holly Ahern is a science writer and an assistant professor of biology at Adirondack Community College in Queensbury, N.Y
 Image via Wikipedia
Image via WikipediaRead more: Sequencing Technologies Helping - The Scientist - Magazine of the Life Sciences http://www.the-scientist.com/article/display/16721/#ixzz1CZTrRIL3

No comments:
Post a Comment
Note: Only a member of this blog may post a comment.